Introduction to Immunoglobulins
Immunoglobulins (Ig), commonly known as antibodies, are glycoprotein molecules produced by B-lymphocytes (B-cells) and plasma cells in response to the presence of antigens. They are essential components of the adaptive immune system, functioning as the humoral arm of immunity. Immunoglobulins are crucial for recognition, neutralization, and elimination of pathogens including bacteria, viruses, toxins, and other foreign substances.
The primary function of immunoglobulins is to specifically bind antigens through highly variable regions, thereby facilitating their removal by immune mechanisms. This specificity is achieved through the variable domains of the immunoglobulin, which form the antigen-binding sites.
Immunoglobulins are soluble proteins present in various body fluids such as blood plasma, lymph, saliva, breast milk, and mucosal secretions, allowing them to perform immune surveillance both systemically and at mucosal surfaces.
Classification of Immunoglobulins
In humans, immunoglobulins are classified into five major classes based on their structure, function, and distribution:
| Class | Abbreviation | Major Function | Location | Special Features |
| IgG | IgG | Long-term immunity, opsonization, complement activation, crosses placenta | Blood, lymph | Most abundant (~75-80% of total Ig), monomeric |
| IgA | IgA | Mucosal immunity, neutralization | Secretions (saliva, tears, mucus, breast milk) | Dimeric in secretions, protects mucosal surfaces |
| IgM | IgM | Primary immune response, complement activation | Blood, lymph | Pentameric, first antibody produced, large size |
| IgE | IgE | Defense against parasites, allergy mediation | Blood, tissues | Monomeric, binds to mast cells and basophils |
| IgD | IgD | B-cell receptor function, antigen recognition | B-cell surfaces, blood | Monomeric, very low concentration |
General Structure of Immunoglobulins
Immunoglobulins share a basic Y-shaped structure composed of four polypeptide chains:
- Two Heavy (H) Chains – longer chains, molecular weight ~50 kDa each.
- Two Light (L) Chains – shorter chains, molecular weight ~25 kDa each.
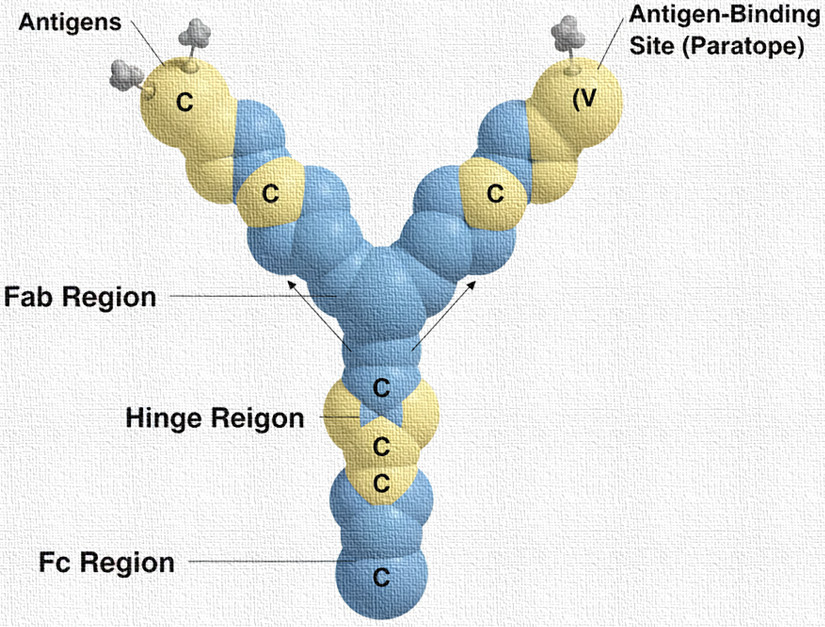
These chains are connected by disulfide bonds, forming a tetrameric structure. The heavy and light chains are further divided into regions:
- Variable (V) region – responsible for antigen binding, located at the N-terminal end of both heavy and light chains. This region exhibits extreme diversity to allow recognition of a vast array of antigens.
- Constant (C) region – responsible for effector functions such as complement activation, opsonization, and binding to immune cells. Found in both light and heavy chains. In heavy chains, the constant region also determines the immunoglobulin class (IgG, IgA, IgM, IgE, IgD).
Detailed Domains
Each polypeptide chain is composed of globular domains stabilized by intra-chain disulfide bonds:
Fab (Fragment, antigen-binding) region
- Located at the arms of the Y.
- Contains the variable domains of both heavy (VH) and light (VL) chains.
- Responsible for specific recognition and binding of antigens.
- Each Ig molecule has two identical Fab regions, allowing bivalent antigen binding.
Fc (Fragment, crystallizable) region
- Forms the stem of the Y.
- Composed of the constant domains of the heavy chains.
- Responsible for effector functions, such as:
- Binding to Fc receptors on phagocytes (opsonization)
- Activation of the complement system
- Mediating antibody-dependent cellular cytotoxicity (ADCC)
Hinge Region
- Flexible segment connecting Fab and Fc regions.
- Provides structural flexibility for simultaneous binding to antigens at different orientations.
Molecular Features
- Glycosylation: Immunoglobulins are glycoproteins, with carbohydrate moieties attached to the Fc region. Glycosylation is crucial for stability, solubility, and effector function.
- Disulfide Bonds: Maintain the structural integrity of the molecule; heavy-light chain disulfide bonds link chains, while intra-chain bonds stabilize the domains.
- Antigen-Binding Site (Paratope): Formed by the hypervariable loops (complementarity-determining regions, CDRs) in the variable regions of heavy and light chains. Each Ig molecule has two identical antigen-binding sites, enabling high specificity.
- Flexibility: The hinge region allows the Fab arms to pivot and reach antigens at varying distances.
Functional Summary
- Neutralization of Pathogens: Antibodies bind to microbial surface molecules or toxins, preventing their interaction with host cells.
- Opsonization: Antibody-coated pathogens are more efficiently recognized and engulfed by phagocytes.
- Complement Activation: Antibodies initiate the classical complement pathway, leading to pathogen lysis.
- Immune Cell Recruitment: Fc regions interact with Fc receptors on NK cells, macrophages, and mast cells to trigger immune responses.
- Antibody Diversity: Achieved through somatic recombination (V(D)J recombination), allowing recognition of millions of different antigens.
Summary of Immunoglobulin Structure
| Feature | Description |
| Shape | Y-shaped, tetrameric structure |
| Chains | 2 Heavy (H) + 2 Light (L) chains |
| Regions | Variable (V) and Constant (C) regions |
| Functional Parts | Fab (antigen-binding), Fc (effector functions), Hinge (flexibility) |
| Bonds | Disulfide bonds (inter- and intra-chain) |
| Glycosylation | Carbohydrate moieties in Fc for stability and function |
| Antigen-binding | Complementarity-determining regions (CDRs) in Fab |

